Lung cancer has the second highest morbidity among all cancers and the highest mortality worldwide according to the latest statistics from the World Health Organization (1,2). Because the disease is often diagnosed at more advanced stages, early detection is essential to increasing lung cancer survival rates (3). Previous work to characterize the tumor microenvironment (TME) and surrounding immune cell populations show lung cancer stage-dependent changes in CD4+ and CD8+ cells, among other cell surface markers (CDs) (4). Methods where CDs are well maintained during processing could enable early detection.
The most clinically relevant methods to research the TME and immune cell populations include flow cytometry and single cell RNA sequencing (scRNA-seq), both of which require upstream preparation of single cell suspensions by tissue dissociation (5). While the conventional methods for generating single cell suspensions have relied on dissociation by heat and enzymes, the Cellsonics SimpleFlow platform uses Bulk Lateral Ultrasonic (BLU) energy for gentle mechanical tissue dissociation, while maintaining samples at a constant 6°C. We have previously demonstrated that this workflow retains more immune cells and immune cell markers for immunophenotyping than dissociation by heat and enzymes. Further, recent work has demonstrated that transcript capture can also be affected by enzymatic tissue dissociation (6), suggesting another advantage of scRNAseq sample preparation with the enzyme and heat-free SimpleFlow BLU processing.
To compare SimpleFlow BLU dissociation to conventional enzymatic dissociation of lung tissue, lungs were harvested from commercially purchased mice (N = 5). Enzyme and heat dissociation was performed with a commercially available mouse lung dissociation kit according to the manufacturer’s protocol and BLU dissociation was preceded only by a manual tissue mincing on ice. Single cell suspensions were then assessed for several cell surface markers and for cell counts using CountBright™ absolute counting beads. Marker expression was determined by FlowJo and two-sample t-tests were performed in GraphPad prism to determine significance of the differences in means between the enzymatic condition and the various BLU conditions.
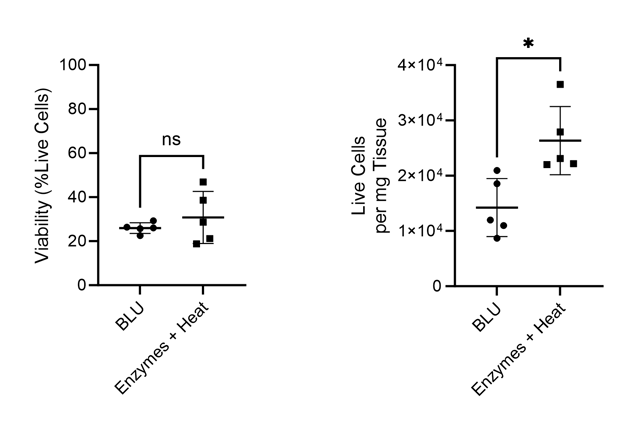
Figure 1. Comparison of cell count per milligram of tissue and total cell viability following dissociation via enzymatic or BLU approaches on lung tissue harvested from commercially purchased mice (N = 5). ns = not significant, * = p-value ≤ 0.05.
While we found that BLU isolated fewer total cells from the lung tissue than dissociation by enzymes and heat, those cells which were isolated had comparable viability (Fig. 1). Interestingly, despite the lower number of overall dissociated cells, the cell surface expression of T cell markers CD3 and CD8 was significantly higher for cells dissociated by BLU, translating into a significantly larger number of CD8+ T cells (Fig. 2).
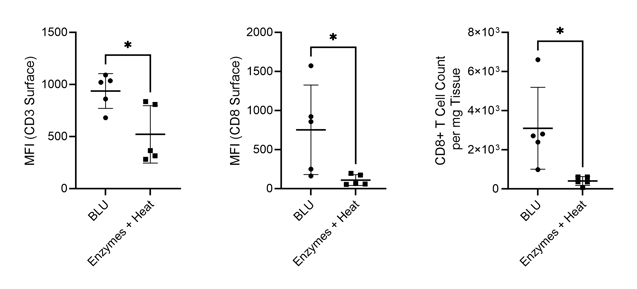
Figure 2. Comparison of CD3 and CD8 expression on live cells and CD8+ T cell count per milligram of tissue isolated via enzymatic or BLU dissociation of lung tissue harvested from commercially purchased mice (N = 5). * = p-value ≤ 0.05.
A closer look at surface markers PD-1 and TIM-3 on CD8+ T cells dissociated both by BLU and by enzymatic approaches (Fig. 3), revealing an interesting and significant loss in median fluorescence intensity (MFI) of both markers on cells dissociated by enzymes and heat. Gating strategies for identifying these populations of interest are shown in Fig. 4.
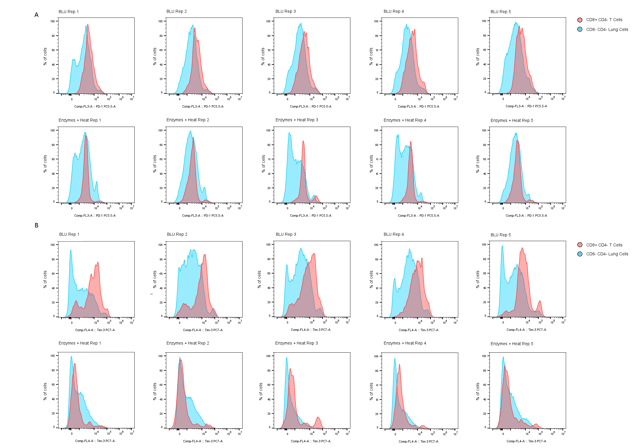
Figure 3. Comparison of (A) PD-1 and (B) TIM-3 expression on CD8+ CD4- T cells (red peaks) and CD8- CD4- lung cells (blue peaks) dissociated by BLU or enzymatic methods (N = 5).
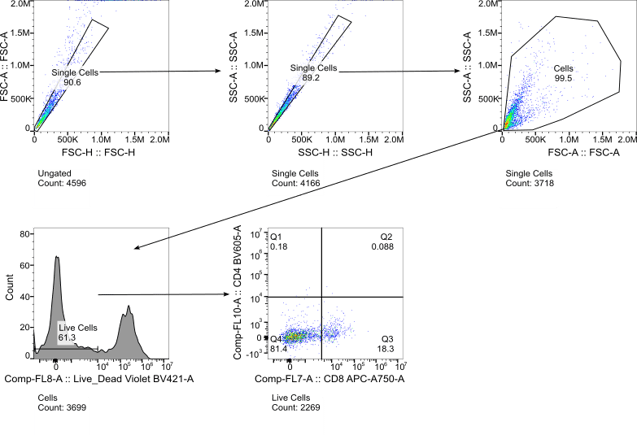
Figure 4. Gating strategy for identifying subsets of lung cells dissociated by both BLU and enzymatic approaches.
Recognizing that both PD-1 and TIM-3 have implications for other immune cell types than just CD8+ T cells, we looked back at PD-1 and TIM-3 expression more generally on all cells dissociated using both methods. Again, we found a significant decrease in expression of both markers on cells dissociated by enzymes and heat (Fig. 5). The artificial downregulation of these markers on all cell populations by enzyme dissociation suggests that cells expressing these markers are either undergoing severe stress, causing them to downregulate these markers, or are being lost or destroyed. Considering that both PD-1 and TIM-3 are markers commonly expressed by cells in situations of stress (7), we suspect that immune cell populations expressing these markers are being lost during methods of enzymatic dissociation but preserved by BLU.
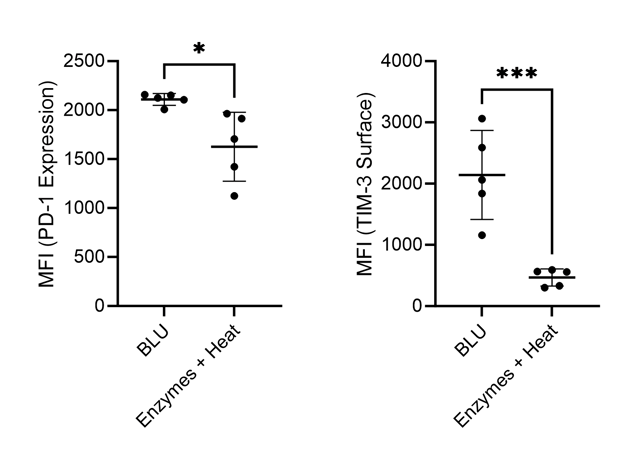
Figure 5. Comparison of PD-1 and TIM-3 expression on lung cells harvested from commercially purchased mice (N = 5). * = p-value ≤ 0.05, *** = p-value ≤ 0.001.
Further, we investigated myeloid cell expression which we have previously demonstrated to be impacted for several cell surface markers by enzymatic dissociation on brain and heart tissue. In cells dissociated by enzymes, we saw a significant decrease in cell surface expression of markers CD24, CD11c, CD11b, and Ly6C (Fig. 6). These artificial losses of expression seen with enzymatic dissociation of lung tissue are highly concerning and suggest that improper conclusions may be drawn from analyses currently employing these methods.
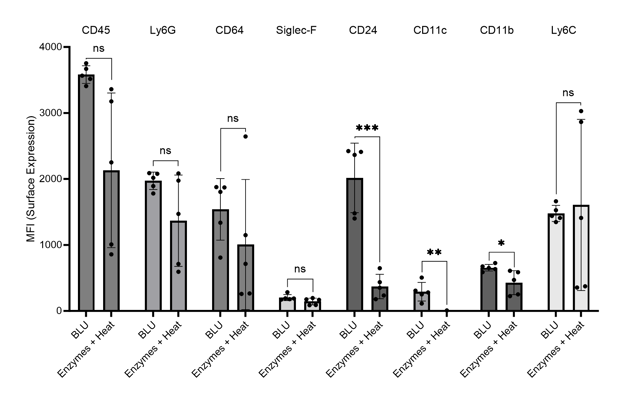
Figure 6. Comparison of myeloid cell marker expression on lung cells harvested from commercially purchased mice (N = 5). * = p-value ≤ 0.05, ** = p-value ≤ 0.01, *** = p-value ≤ 0.001.
As an alternative, BLU dissociation preserves these markers and enables a higher fidelity single cell collection for downstream processing. While we will continue to enhance protocols for the dissociation of lung and other tissue types, we recommend that researchers wanting to obtain high quality and reliable single cell suspensions from their tissue samples consider switching to SimpleFlow.
References
- Cancer [Internet]. [cited 2023 Aug 21]. Available from: https://www.who.int/news-room/fact-sheets/detail/cancer
- Yuan M, Zhao Y, Arkenau HT, Lao T, Chu L, Xu Q. Signal pathways and precision therapy of small-cell lung cancer. Signal Transduct Target Ther. 2022 Jun 15;7(1):1–18.
- Nana-Sinkam SP, Powell CA. Molecular Biology of Lung Cancer: Diagnosis and Management of Lung Cancer, 3rd ed: American College of Chest Physicians Evidence-Based Clinical Practice Guidelines. CHEST. 2013 May 1;143(5):e30S-e39S.
- Immune and Inflammatory Cell Composition of Human Lung Cancer Stroma | PLOS ONE [Internet]. [cited 2023 Aug 21]. Available from: https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0139073
- Olesch C, Brunn D, Aktay-Cetin Ö, Sirait-Fischer E, Pullamsetti SS, Grimminger F, et al. Picturing of the Lung Tumor Cellular Composition by Multispectral Flow Cytometry. Front Immunol [Internet]. 2022 [cited 2023 Aug 21];13. Available from: https://www.frontiersin.org/articles/10.3389/fimmu.2022.827719
- Quintanal-Villalonga Á, Chan JM, Masilionis I, Gao VR, Xie Y, Allaj V, et al. Protocol to dissociate, process, and analyze the human lung tissue using single-cell RNA-seq. STAR Protoc. 2022 Dec;3(4):101776.
- Xia Q, Wei L, Zhang Y, Sheng J, Wu W, Zhang Y. Immune Checkpoint Receptors Tim-3 and PD-1 Regulate Monocyte and T Lymphocyte Function in Septic Patients. Mediators Inflamm. 2018 Nov 22;2018:e1632902.
Interested in learning more?
Ask for a 20-minute seminar from a Cellsonics Scientist




.png?width=300&height=54&name=CellsonicsLogoWhite(2048%20x%20366%20px).png)